Tutorial 3: Simultaneous fitting/regression#
Week 3, Day 5: Network Causality
By Neuromatch Academy
Content creators: Ari Benjamin, Tony Liu, Konrad Kording
Content reviewers: Mike X Cohen, Madineh Sarvestani, Yoni Friedman, Ella Batty, Michael Waskom
Production editors: Gagana B, Spiros Chavlis
Tutorial objectives#
Estimated timing of tutorial: 20 min
This is tutorial 3 on our day of examining causality. Below is the high level outline of what we’ll cover today, with the sections we will focus on in this notebook in bold:
Master definitions of causality
Understand that estimating causality is possible
Learn 4 different methods and understand when they fail
perturbations
correlations
simultaneous fitting/regression
instrumental variables
Tutorial 3 objectives
In tutorial 2 we explored correlation as an approximation for causation and learned that correlation \(\neq\) causation for larger networks. However, computing correlations is a rather simple approach, and you may be wondering: will more sophisticated techniques allow us to better estimate causality? Can’t we control things?
Here we’ll use some common advanced (but controversial) methods that estimate causality from observational data. These methods rely on fitting a function to our data directly, instead of trying to use perturbations or correlations. Since we have the full closed-form equation of our system, we can try these methods and see how well they work in estimating causal connectivity when there are no perturbations. Specifically, we will:
Learn about more advanced (but also controversial) techniques for estimating causality
conditional probabilities (regression)
Explore limitations and failure modes
understand the problem of omitted variable bias
Setup#
Install and import feedback gadget#
Show code cell source
# @title Install and import feedback gadget
!pip3 install vibecheck datatops --quiet
from vibecheck import DatatopsContentReviewContainer
def content_review(notebook_section: str):
return DatatopsContentReviewContainer(
"", # No text prompt
notebook_section,
{
"url": "https://pmyvdlilci.execute-api.us-east-1.amazonaws.com/klab",
"name": "neuromatch_cn",
"user_key": "y1x3mpx5",
},
).render()
feedback_prefix = "W3D5_T3"
# Imports
import numpy as np
import matplotlib.pyplot as plt
from sklearn.multioutput import MultiOutputRegressor
from sklearn.linear_model import Lasso
Figure Settings#
Show code cell source
# @title Figure Settings
import logging
logging.getLogger('matplotlib.font_manager').disabled = True
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/main/nma.mplstyle")
Plotting Functions#
Show code cell source
# @title Plotting Functions
def see_neurons(A, ax, ratio_observed=1, arrows=True, show=False):
"""
Visualizes the connectivity matrix.
Args:
A (np.ndarray): the connectivity matrix of shape (n_neurons, n_neurons)
ax (plt.axis): the matplotlib axis to display on
Returns:
Nothing, but visualizes A.
"""
n = len(A)
ax.set_aspect('equal')
thetas = np.linspace(0, np.pi * 2, n, endpoint=False)
x, y = np.cos(thetas), np.sin(thetas),
if arrows:
for i in range(n):
for j in range(n):
if A[i, j] > 0:
ax.arrow(x[i], y[i], x[j] - x[i], y[j] - y[i], color='k',
head_width=.05, width = A[i, j] / 25, shape='right',
length_includes_head=True, alpha=.2)
if ratio_observed < 1:
nn = int(n * ratio_observed)
ax.scatter(x[:nn], y[:nn], c='r', s=150, label='Observed')
ax.scatter(x[nn:], y[nn:], c='b', s=150, label='Unobserved')
ax.legend(fontsize=15)
else:
ax.scatter(x, y, c='k', s=150)
ax.axis('off')
if show:
plt.show()
def plot_connectivity_matrix(A, ax=None):
"""Plot the (weighted) connectivity matrix A as a heatmap
Args:
A (ndarray): connectivity matrix (n_neurons by n_neurons)
ax: axis on which to display connectivity matrix
"""
if ax is None:
ax = plt.gca()
lim = np.abs(A).max()
ax.imshow(A, vmin=-lim, vmax=lim, cmap="coolwarm")
plt.show()
Helper Functions#
Show code cell source
# @title Helper Functions
def sigmoid(x):
"""
Compute sigmoid nonlinearity element-wise on x.
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with sigmoid nonlinearity applied
"""
return 1 / (1 + np.exp(-x))
def create_connectivity(n_neurons, random_state=42, p=0.9):
"""
Generate our nxn causal connectivity matrix.
Args:
n_neurons (int): the number of neurons in our system.
random_state (int): random seed for reproducibility
Returns:
A (np.ndarray): our 0.1 sparse connectivity matrix
"""
np.random.seed(random_state)
A_0 = np.random.choice([0, 1], size=(n_neurons, n_neurons), p=[p, 1 - p])
# set the timescale of the dynamical system to about 100 steps
_, s_vals, _ = np.linalg.svd(A_0)
if s_vals[0] != 0 and not np.isnan(s_vals[0]):
A = A_0 / (1.01 * s_vals[0])
else:
eps = 1e-12
A = eps*np.ones_like(A_0) # if denominator is zero, set A to a small value
return A
def simulate_neurons(A, timesteps, random_state=42):
"""
Simulates a dynamical system for the specified number of neurons and timesteps.
Args:
A (np.array): the connectivity matrix
timesteps (int): the number of timesteps to simulate our system.
random_state (int): random seed for reproducibility
Returns:
X has shape (n_neurons, timeteps).
"""
np.random.seed(random_state)
n_neurons = len(A)
X = np.zeros((n_neurons, timesteps))
for t in range(timesteps - 1):
epsilon = np.random.multivariate_normal(np.zeros(n_neurons),
np.eye(n_neurons))
X[:, t + 1] = sigmoid(A.dot(X[:, t]) + epsilon)
assert epsilon.shape == (n_neurons, )
return X
def get_sys_corr(n_neurons, timesteps, random_state=42, neuron_idx=None):
"""
A wrapper function for our correlation calculations between A and R.
Args:
n_neurons (int): the number of neurons in our system.
timesteps (int): the number of timesteps to simulate our system.
random_state (int): seed for reproducibility
neuron_idx (int): optionally provide a neuron idx to slice out
Returns:
A single float correlation value representing the similarity between A and R
"""
A = create_connectivity(n_neurons, random_state)
X = simulate_neurons(A, timesteps)
R = correlation_for_all_neurons(X)
return np.corrcoef(A.flatten(), R.flatten())[0, 1]
def correlation_for_all_neurons(X):
"""
Computes the connectivity matrix for the all neurons using correlations
Args:
X: the matrix of activities
Returns:
estimated_connectivity (np.ndarray): estimated connectivity for the
selected neuron, of shape (n_neurons,)
"""
n_neurons = len(X)
S = np.concatenate([X[:, 1:], X[:, :-1]], axis=0)
R = np.corrcoef(S)[:n_neurons, n_neurons:]
return R
The helper functions defined above are:
sigmoid: computes sigmoid nonlinearity element-wise on input, from Tutorial 1create_connectivity: generates nxn causal connectivity matrix., from Tutorial 1simulate_neurons: simulates a dynamical system for the specified number of neurons and timesteps, from Tutorial 1get_sys_corr: a wrapper function for correlation calculations between A and R, from Tutorial 2correlation_for_all_neurons: computes the connectivity matrix for the all neurons using correlations, from Tutorial 2
Section 1: Regression: recovering connectivity by model fitting#
Video 1: Regression approach#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Regression_approach_Video")
You may be familiar with the idea that correlation only implies causation when there are no hidden confounders. This aligns with our intuition that correlation only implies causality when no alternative variables could explain away a correlation.
A confounding example: Suppose you observe that people who sleep more do better in school. It’s a nice correlation. But what else could explain it? Maybe people who sleep more are richer, don’t work a second job, and have time to actually do homework. If you want to ask if sleep causes better grades, and want to answer that with correlations, you have to control for all possible confounds.
A confound is any variable that affects both the outcome and your original covariate. In our example, confounds are things that affect both sleep and grades.
Controlling for a confound: Confonds can be controlled for by adding them as covariates in a regression. But for your coefficients to be causal effects, you need three things:
All confounds are included as covariates
Your regression assumes the same mathematical form of how covariates relate to outcomes (linear, GLM, etc.)
No covariates are caused by both the treatment (original variable) and the outcome. These are colliders; we won’t introduce it today (but Google it on your own time! Colliders are very counterintuitive.)
In the real world it is very hard to guarantee these conditions are met. In the brain it’s even harder (as we can’t measure all neurons). Luckily today we simulated the system ourselves.
Video 2: Fitting a GLM#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Fitting_a_GLM_Video")
Recall that in our system each neuron effects every other via:
where \(\sigma\) is our sigmoid nonlinearity from before: \(\sigma(x) = \frac{1}{1 + e^{-x}}\)
Our system is a closed system, too, so there are no omitted variables. The regression coefficients should be the causal effect. Are they?
We will use a regression approach to estimate the causal influence of all neurons to neuron #1. Specifically, we will use linear regression to determine the \(A\) in:
where \(\sigma^{-1}\) is the inverse sigmoid transformation, also sometimes referred to as the logit transformation: \(\sigma^{-1}(x) = \log(\frac{x}{1-x})\).
Let \(W\) be the \(\vec{x}_t\) values, up to the second-to-last timestep \(T-1\):
Let \(Y\) be the \(\vec{x}_{t+1}\) values for a selected neuron, indexed by \(i\), starting from the second timestep up to the last timestep \(T\):
You will then fit the following model:
where \(V\) is the \(n \times 1\) coefficient matrix of this regression, which will be the estimated connectivity matrix between the selected neuron and the rest of the neurons.
Review: As you learned in Week 1, lasso a.k.a. \(L_1\) regularization causes the coefficients to be sparse, containing mostly zeros. Think about why we want this here.
Coding Exercise 1: Use linear regression plus lasso to estimate causal connectivities#
You will now create a function to fit the above regression model and V. We will then call this function to examine how close the regression vs the correlation is to true causality.
Code:
You’ll notice that we’ve transposed both \(Y\) and \(W\) here and in the code we’ve already provided below. Why is that?
This is because the machine learning models provided in scikit-learn expect the rows of the input data to be the observations, while the columns are the variables. We have that inverted in our definitions of \(Y\) and \(W\), with the timesteps of our system (the observations) as the columns. So we transpose both matrices to make the matrix orientation correct for scikit-learn.
Because of the abstraction provided by scikit-learn, fitting this regression will just be a call to initialize the
Lasso()estimator and a call to thefit()functionUse the following hyperparameters for the
Lassoestimator:alpha = 0.01fit_intercept = False
How do we obtain \(V\) from the fitted model?
We will use the helper function logit.
# Set parameters
n_neurons = 50 # the size of our system
timesteps = 10000 # the number of timesteps to take
random_state = 42
neuron_idx = 1
# Set up system and simulate
A = create_connectivity(n_neurons, random_state)
X = simulate_neurons(A, timesteps)
Execute this cell to enable helper function logit
Show code cell source
# @markdown Execute this cell to enable helper function `logit`
def logit(x):
"""
Applies the logit (inverse sigmoid) transformation
Args:
x (np.ndarray): the numpy data array we want to transform
Returns
(np.ndarray): x with logit nonlinearity applied
"""
return np.log(x/(1-x))
def get_regression_estimate(X, neuron_idx):
"""
Estimates the connectivity matrix using lasso regression.
Args:
X (np.ndarray): our simulated system of shape (n_neurons, timesteps)
neuron_idx (int): a neuron index to compute connectivity for
Returns:
V (np.ndarray): estimated connectivity matrix of shape (n_neurons, n_neurons).
if neuron_idx is specified, V is of shape (n_neurons,).
"""
# Extract Y and W as defined above
W = X[:, :-1].transpose()
Y = X[[neuron_idx], 1:].transpose()
# Apply inverse sigmoid transformation
Y = logit(Y)
############################################################################
## TODO: Insert your code here to fit a regressor with Lasso. Lasso captures
## our assumption that most connections are precisely 0.
## Fill in function and remove
raise NotImplementedError("Please complete the regression exercise")
############################################################################
# Initialize regression model with no intercept and alpha=0.01
regression = ...
# Fit regression to the data
regression.fit(...)
V = regression.coef_
return V
# Estimate causality with regression
V = get_regression_estimate(X, neuron_idx)
print(f"Regression: correlation of estimated with true connectivity: {np.corrcoef(A[neuron_idx, :], V)[1, 0]:.3f}")
print(f"Lagged correlation of estimated with true connectivity: {get_sys_corr(n_neurons, timesteps, random_state, neuron_idx=neuron_idx):.3f}")
You should find that:
Regression: correlation of estimated connectivity with true connectivity: 0.865
Lagged correlation of estimated connectivity with true connectivity: 0.703
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Linear_regression_with_lasso_to_estimate_causal_connectivities_Exercise")
We can see from these numbers that multiple regression is better than simple correlation for estimating connectivity.
Section 2: Partially Observed Systems#
Estimated timing to here from start of tutorial: 10 min
If we are unable to observe the entire system, omitted variable bias becomes a problem. If we don’t have access to all the neurons, and so therefore can’t control them, can we still estimate the causal effect accurately?
Video 3: Omitted variable bias#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Omitted_variable_bias_Video")
Video correction: the labels “connectivity from”/”connectivity to” are swapped in the video but fixed in the figures/demos below
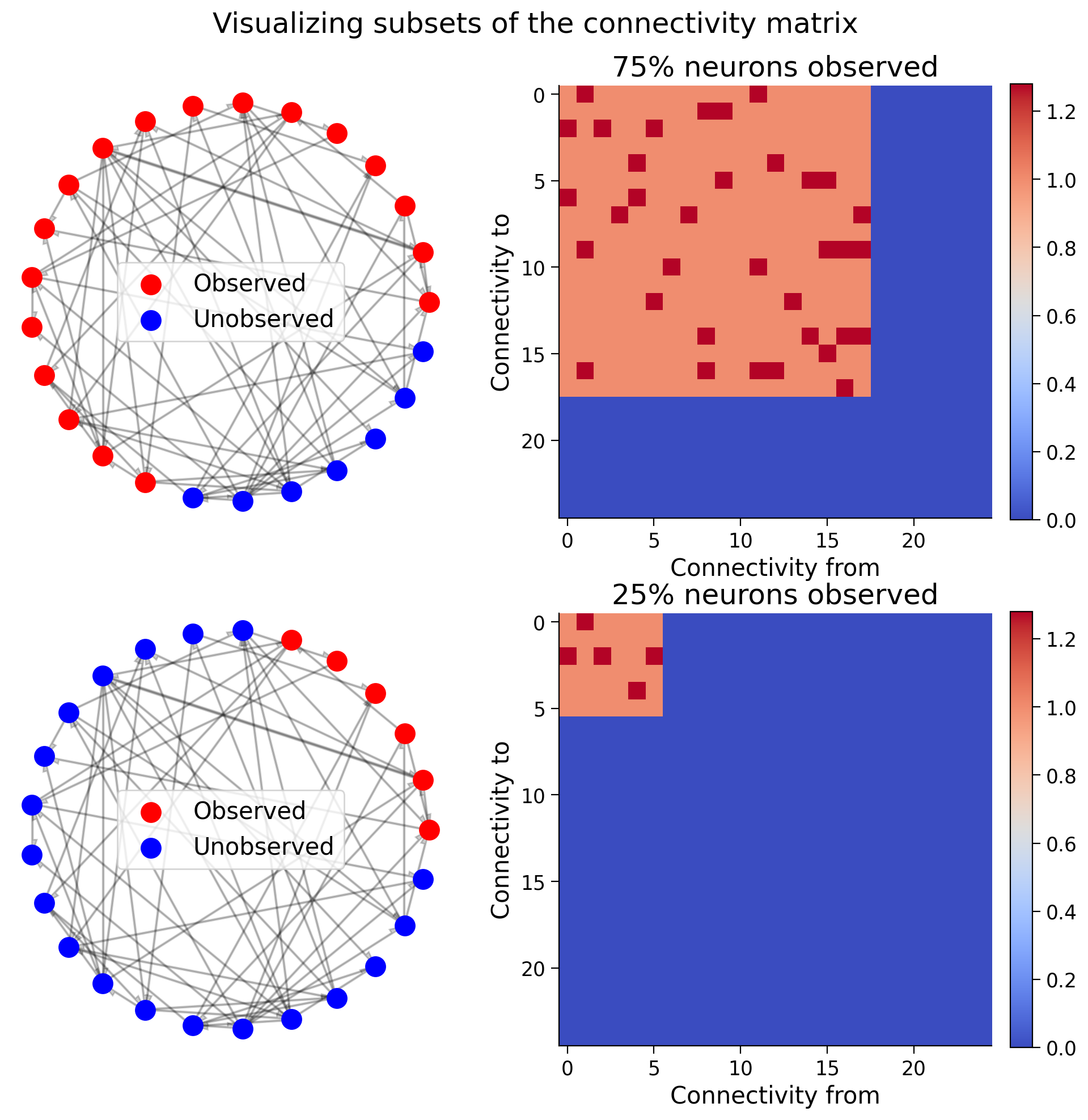
We first visualize different subsets of the connectivity matrix when we observe 75% of the neurons vs 25%.
Recall the meaning of entries in our connectivity matrix: \(A[i,j] = 1\) means a connectivity from neuron \(i\) to neuron \(j\) with strength \(1\).
Execute this cell to visualize subsets of connectivity matrix
Show code cell source
# @markdown Execute this cell to visualize subsets of connectivity matrix
# Run this cell to visualize the subsets of variables we observe
n_neurons = 25
A = create_connectivity(n_neurons)
fig, axs = plt.subplots(2, 2, figsize=(10, 10))
ratio_observed = [0.75, 0.25] # the proportion of neurons observed in our system
for i, ratio in enumerate(ratio_observed):
sel_idx = int(n_neurons * ratio)
offset = np.zeros((n_neurons, n_neurons))
axs[i, 1].title.set_text(f"{int(ratio * 100)}% neurons observed")
offset[:sel_idx, :sel_idx] = 1 + A[:sel_idx, :sel_idx]
im = axs[i, 1].imshow(offset, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
axs[i, 1].set_xlabel("Connectivity from")
axs[i, 1].set_ylabel("Connectivity to")
plt.colorbar(im, ax=axs[i, 1], fraction=0.046, pad=0.04)
see_neurons(A, axs[i, 0], ratio)
plt.suptitle("Visualizing subsets of the connectivity matrix")
plt.tight_layout()
plt.show()
Interactive Demo 3: Regression performance as a function of the number of observed neurons#
We will first change the number of observed neurons in the network and inspect the resulting estimates of connectivity in this interactive demo. How does the estimated connectivity differ?
Execute this cell to get helper functions get_regression_estimate_full_connectivity and get_regression_corr_full_connectivity
Show code cell source
# @markdown Execute this cell to get helper functions `get_regression_estimate_full_connectivity` and `get_regression_corr_full_connectivity`
def get_regression_estimate_full_connectivity(X):
"""
Estimates the connectivity matrix using lasso regression.
Args:
X (np.ndarray): our simulated system of shape (n_neurons, timesteps)
neuron_idx (int): optionally provide a neuron idx to compute connectivity for
Returns:
V (np.ndarray): estimated connectivity matrix of shape (n_neurons, n_neurons).
if neuron_idx is specified, V is of shape (n_neurons,).
"""
n_neurons = X.shape[0]
# Extract Y and W as defined above
W = X[:, :-1].transpose()
Y = X[:, 1:].transpose()
# apply inverse sigmoid transformation
Y = logit(Y)
# fit multioutput regression
reg = MultiOutputRegressor(Lasso(fit_intercept=False,
alpha=0.01, max_iter=250),
n_jobs=-1)
reg.fit(W, Y)
V = np.zeros((n_neurons, n_neurons))
for i, estimator in enumerate(reg.estimators_):
V[i, :] = estimator.coef_
return V
def get_regression_corr_full_connectivity(n_neurons, A, X, observed_ratio,
regression_args):
"""
A wrapper function for our correlation calculations between A and the V estimated
from regression.
Args:
n_neurons (int): number of neurons
A (np.ndarray): connectivity matrix
X (np.ndarray): dynamical system
observed_ratio (float): the proportion of n_neurons observed, must be between 0 and 1.
regression_args (dict): dictionary of lasso regression arguments and hyperparameters
Returns:
A single float correlation value representing the similarity between A and R
"""
assert (observed_ratio > 0) and (observed_ratio <= 1)
sel_idx = np.clip(int(n_neurons*observed_ratio), 1, n_neurons)
sel_X = X[:sel_idx, :]
sel_A = A[:sel_idx, :sel_idx]
sel_V = get_regression_estimate_full_connectivity(sel_X)
return np.corrcoef(sel_A.flatten(), sel_V.flatten())[1, 0], sel_V
Show code cell source
# @markdown Execute this cell to enable the widgets
# @markdown Note: The plots will take a few seconds to update after moving the slider.
n_neurons = 50
A = create_connectivity(n_neurons, random_state=42)
X = simulate_neurons(A, 4000, random_state=42)
reg_args = {
"fit_intercept": False,
"alpha": 0.001
}
@widgets.interact(n_observed=widgets.IntSlider(min=5, max=45, step=5,
continuous_update=False))
def plot_observed(n_observed):
to_neuron = 0
fig, axs = plt.subplots(1, 3, figsize=(15, 5))
sel_idx = n_observed
ratio = (n_observed) / n_neurons
offset = np.zeros((n_neurons, n_neurons))
axs[0].title.set_text(f"{int(ratio * 100)}% neurons observed")
offset[:sel_idx, :sel_idx] = 1 + A[:sel_idx, :sel_idx]
im = axs[1].imshow(offset, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
plt.colorbar(im, ax=axs[1], fraction=0.046, pad=0.04)
see_neurons(A,axs[0], ratio, False)
corr, R = get_regression_corr_full_connectivity(n_neurons, A, X,
ratio, reg_args)
big_R = np.zeros(A.shape)
big_R[:sel_idx, :sel_idx] = 1 + R
im = axs[2].imshow(big_R, cmap="coolwarm", vmin=0, vmax=A.max() + 1)
plt.colorbar(im, ax=axs[2],fraction=0.046, pad=0.04)
c = 'w' if n_observed<(n_neurons-3) else 'k'
axs[2].text(0, n_observed + 3, f"Correlation: {corr:.2f}", color=c, size=15)
axs[1].title.set_text("True connectivity")
axs[1].set_xlabel("Connectivity from")
axs[1].set_ylabel("Connectivity to")
axs[2].title.set_text("Estimated connectivity")
axs[2].set_xlabel("Connectivity from")
plt.show()
Next, we will inspect a plot of the correlation between true and estimated connectivity matrices vs the percent of neurons observed over multiple trials. What is the relationship that you see between performance and the number of neurons observed?
Note: the cell below will take about 25-30 seconds to run.
Plot correlation vs. subsampling
Show code cell source
# @markdown Plot correlation vs. subsampling
# we'll simulate many systems for various ratios of observed neurons
n_neurons = 50
timesteps = 5000
ratio_observed = [1, 0.75, 0.5, .25, .125] # the proportion of neurons observed in our system
n_trials = 3 # run it this many times to get variability in our results
reg_args = {
"fit_intercept": False,
"alpha": 0.001
}
corr_data = np.zeros((n_trials, len(ratio_observed)))
for trial in range(n_trials):
A = create_connectivity(n_neurons, random_state=trial)
X = simulate_neurons(A, timesteps)
print(f"simulating trial {trial + 1} of {n_trials}")
for j, ratio in enumerate(ratio_observed):
result,_ = get_regression_corr_full_connectivity(n_neurons, A, X,
ratio, reg_args)
corr_data[trial, j] = result
corr_mean = np.nanmean(corr_data, axis=0)
corr_std = np.nanstd(corr_data, axis=0)
plt.plot(np.asarray(ratio_observed) * 100, corr_mean)
plt.fill_between(np.asarray(ratio_observed) * 100,
corr_mean - corr_std, corr_mean + corr_std,
alpha=.2)
plt.xlim([100, 10])
plt.xlabel("Percent of neurons observed")
plt.ylabel("connectivity matrices correlation")
plt.title("Performance of regression\nas a function of the number of neurons observed")
plt.show()
simulating trial 1 of 3
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.035e-01, tolerance: 6.170e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.194e+00, tolerance: 5.991e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.444e+01, tolerance: 1.037e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.527e+00, tolerance: 7.297e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 5.612e+01, tolerance: 8.502e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.267e-01, tolerance: 6.659e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.571e+00, tolerance: 7.069e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.784e+00, tolerance: 7.325e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.205e+00, tolerance: 9.716e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.105e+00, tolerance: 6.815e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.791e+00, tolerance: 7.903e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.111e+00, tolerance: 5.913e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.089e-01, tolerance: 6.599e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.078e+00, tolerance: 6.658e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.302e+00, tolerance: 7.114e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.171e+00, tolerance: 7.299e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.018e+00, tolerance: 6.731e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.472e+00, tolerance: 7.109e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.636e+00, tolerance: 7.693e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.060e+01, tolerance: 7.243e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.154e+00, tolerance: 6.479e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.467e+00, tolerance: 7.440e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.440e+00, tolerance: 7.260e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.721e+00, tolerance: 6.617e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.637e-01, tolerance: 6.640e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.633e+00, tolerance: 7.605e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.259e+01, tolerance: 8.183e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.021e-01, tolerance: 5.991e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.204e-01, tolerance: 6.170e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.223e+00, tolerance: 1.037e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.172e+00, tolerance: 7.297e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.555e+01, tolerance: 8.502e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.058e+00, tolerance: 6.659e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.163e+00, tolerance: 7.325e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.365e+00, tolerance: 7.069e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.059e+00, tolerance: 9.716e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.291e+00, tolerance: 6.815e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.818e+00, tolerance: 7.903e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.122e+00, tolerance: 6.599e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.162e+00, tolerance: 6.658e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.352e+00, tolerance: 7.114e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.268e+00, tolerance: 7.299e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.813e+01, tolerance: 7.243e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.262e-01, tolerance: 7.260e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.152e+00, tolerance: 6.731e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.719e+00, tolerance: 7.693e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.390e-01, tolerance: 7.440e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.202e+00, tolerance: 6.617e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.340e-01, tolerance: 6.522e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.547e+00, tolerance: 7.109e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.700e-01, tolerance: 6.170e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.904e+00, tolerance: 1.037e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.419e+00, tolerance: 7.297e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.223e+00, tolerance: 6.659e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.428e+00, tolerance: 8.502e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.013e+00, tolerance: 9.716e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.687e+00, tolerance: 7.069e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.523e+00, tolerance: 6.815e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.683e+00, tolerance: 7.903e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.236e+00, tolerance: 7.114e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.665e+00, tolerance: 6.658e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.632e+00, tolerance: 7.299e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.873e+00, tolerance: 1.037e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.514e+00, tolerance: 8.502e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.991e+00, tolerance: 9.716e-01
model = cd_fast.enet_coordinate_descent(
simulating trial 2 of 3
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.731e+00, tolerance: 7.361e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.023e-01, tolerance: 6.741e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.002e+01, tolerance: 8.649e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.238e+00, tolerance: 6.406e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.129e-01, tolerance: 6.262e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.260e-01, tolerance: 6.778e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.617e+00, tolerance: 6.656e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.394e+00, tolerance: 9.632e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.720e-01, tolerance: 5.902e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.908e+00, tolerance: 7.739e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.495e+00, tolerance: 7.708e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.896e+00, tolerance: 7.932e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.209e+00, tolerance: 6.617e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.575e+01, tolerance: 8.446e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.991e+00, tolerance: 7.790e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.312e+01, tolerance: 8.605e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.333e+01, tolerance: 1.165e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.719e-01, tolerance: 5.862e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.998e-01, tolerance: 6.702e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.094e-01, tolerance: 6.741e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.758e+00, tolerance: 7.361e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.478e-01, tolerance: 6.406e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 5.846e+00, tolerance: 8.649e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.533e+00, tolerance: 6.778e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.073e+00, tolerance: 6.656e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.124e-01, tolerance: 6.149e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.776e+00, tolerance: 9.632e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.977e-01, tolerance: 6.159e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.171e+00, tolerance: 7.739e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.102e+00, tolerance: 6.617e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.217e-01, tolerance: 5.514e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.001e+00, tolerance: 8.605e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.723e+00, tolerance: 7.708e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.556e+00, tolerance: 7.932e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.035e+00, tolerance: 6.173e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.325e+01, tolerance: 8.446e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.535e-01, tolerance: 6.413e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.232e+01, tolerance: 1.165e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.339e+00, tolerance: 7.361e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.226e+00, tolerance: 8.649e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.151e+00, tolerance: 6.262e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.598e+00, tolerance: 6.778e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.910e-01, tolerance: 6.656e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.720e-01, tolerance: 6.253e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.940e+00, tolerance: 9.632e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.304e-01, tolerance: 7.739e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.768e+00, tolerance: 8.649e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.679e+00, tolerance: 9.632e-01
model = cd_fast.enet_coordinate_descent(
simulating trial 3 of 3
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.731e-01, tolerance: 6.676e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.265e+00, tolerance: 6.124e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.043e+00, tolerance: 6.539e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.512e+00, tolerance: 9.272e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.578e+01, tolerance: 7.448e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.497e+00, tolerance: 8.507e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.355e+01, tolerance: 1.073e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.344e+00, tolerance: 8.328e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.221e-01, tolerance: 6.339e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.883e+01, tolerance: 1.060e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.189e+01, tolerance: 8.865e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.905e+01, tolerance: 7.096e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.500e-01, tolerance: 6.317e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.462e+00, tolerance: 7.751e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.015e+00, tolerance: 7.284e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.263e-01, tolerance: 6.171e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.905e+00, tolerance: 7.797e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.577e-01, tolerance: 5.916e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.973e+00, tolerance: 7.764e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.120e+00, tolerance: 7.435e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.278e+00, tolerance: 6.878e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.432e+00, tolerance: 7.365e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.042e+00, tolerance: 7.395e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.468e-01, tolerance: 6.390e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.049e+00, tolerance: 6.547e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.916e-01, tolerance: 6.178e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 8.255e-01, tolerance: 6.124e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.204e-01, tolerance: 6.539e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.364e+00, tolerance: 9.272e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 7.252e+00, tolerance: 7.448e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.674e+00, tolerance: 8.507e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 6.873e+00, tolerance: 1.073e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.965e+00, tolerance: 8.328e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.029e+00, tolerance: 6.339e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 5.834e+01, tolerance: 1.060e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.714e+00, tolerance: 8.865e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.032e+00, tolerance: 7.096e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.837e+00, tolerance: 7.751e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.367e+00, tolerance: 7.797e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.713e+00, tolerance: 7.284e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.221e+00, tolerance: 7.435e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 9.202e-01, tolerance: 6.539e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 2.303e+00, tolerance: 9.272e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.476e+00, tolerance: 8.507e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.863e+00, tolerance: 7.448e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 4.466e+00, tolerance: 1.073e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.737e+00, tolerance: 8.328e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 3.224e+00, tolerance: 8.865e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.130e+00, tolerance: 7.096e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 5.684e+00, tolerance: 1.060e+00
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/sklearn/linear_model/_coordinate_descent.py:695: ConvergenceWarning: Objective did not converge. You might want to increase the number of iterations, check the scale of the features or consider increasing regularisation. Duality gap: 1.415e+00, tolerance: 7.751e-01
model = cd_fast.enet_coordinate_descent(
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/numpy/lib/_function_base_impl.py:2922: RuntimeWarning: invalid value encountered in divide
c /= stddev[:, None]
/opt/hostedtoolcache/Python/3.9.25/x64/lib/python3.9/site-packages/numpy/lib/_function_base_impl.py:2923: RuntimeWarning: invalid value encountered in divide
c /= stddev[None, :]
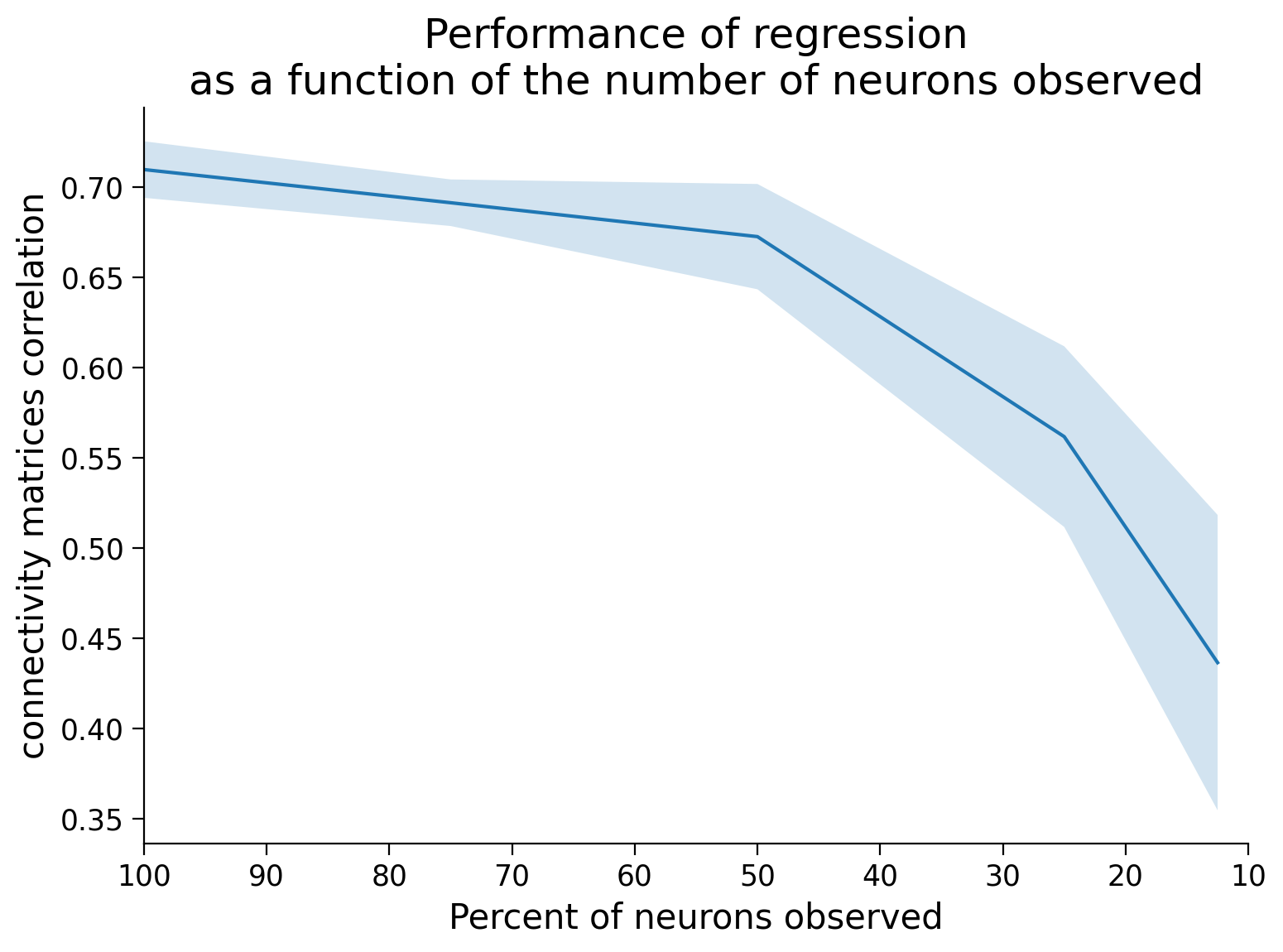
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Regression_performance_as_a_function_of_the_number_of_observed_neurons_Interactive_Demo")
Summary#
Estimated timing of tutorial: 20 min
Video 4: Summary#
Submit your feedback#
Show code cell source
# @title Submit your feedback
content_review(f"{feedback_prefix}_Summary")
In this tutorial, we learned:
To use regression for estimating causality
The problem of omitted variable bias, and how it arises in practice

